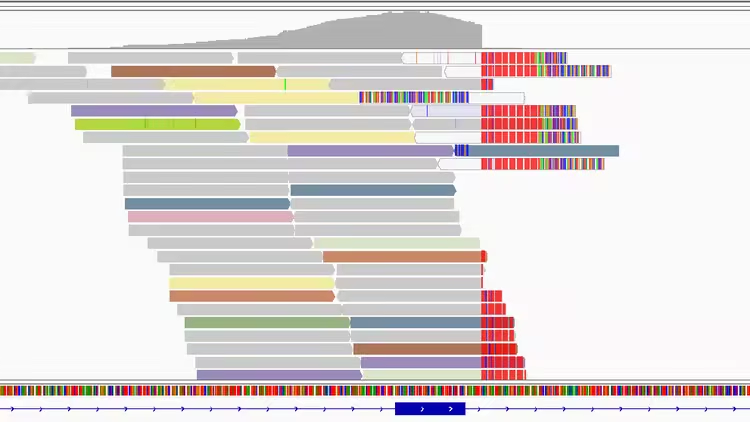
 Free Download Reading structural variants in IGV (short-read NGS)
Free Download Reading structural variants in IGV (short-read NGS)Published 4/2026
MP4 |
Video: h264, 1920x1080 |
Audio: AAC, 44.1 KHz, 2 Ch
Language: English |
Duration: 1h 18m |
Size: 243.96 MB
Learn to visualize and interpret structural variants in IGV using short-read NGS data and real sequencing examples
Recognize the main signals of structural variants in short-read NGS data, including changes in read depth, allele balance, and discordant read pairs.
Visualize deletions, duplications, inversions, insertions, and translocations in the Integrative Genomics Viewer (IGV).
Interpret structural variant patterns in IGV and distinguish true genomic rearrangements from sequencing or alignment artifacts.
Understand the biological and clinical consequences of structural variants, including haploinsufficiency and gene dosage effects.
Apply IGV visualization to support structural variant interpretation in research or clinical genomics.
RequirementsThe course focuses on the interpretation stage of NGS analysis and is intended for those who are already familiar with sequencing data and want to learn how to evaluate, visualize, and confidently interpret CNVs using IGV.
This course assumes that you have a working knowledge of
Basic human genetics and genomics
General principles of next-generation sequencing (NGS)
Common types of genetic variation, including single-nucleotide variants and small indels
No advanced bioinformatics or programming skills are required, and the course does not cover raw NGS data processing or variant calling. Students should have IGV (Integrative Genomics Viewer) installed on their computer to follow the practical modules.
DescriptionCopy number variants and other structural variants play an important role in human genetics and are frequently associated with both normal genomic variation and disease. However, detecting and interpreting these variants can be challenging, particularly when working with short-read next-generation sequencing (NGS) data.
This course provides a practical introduction to recognizing and interpreting structural variants using the Integrative Genomics Viewer (IGV). It is designed for biologists, clinical genomic analysts, and researchers who want to build confidence in working with NGS data and structural variation.
Throughout the course, you will learn how different structural variants appear in short-read sequencing data and how to recognize their characteristic patterns in IGV. Using real examples, we will explore how to identify signatures of deletions, duplications, inversions, translocations, and other structural events, and how these patterns can support variant interpretation. The course focuses on practical pattern recognition and aims to help you develop a more intuitive understanding of how structural variation appears in sequencing data.
By the end of this course, you will be able to confidently navigate NGS data in IGV, recognize common structural variant signatures, and use visualization as a practical tool for interpreting genomic variation.
Whether you are beginning to explore structural variants or looking to strengthen your analytical skills, this course offers clear, hands-on training tailored for professionals working with genomic data.
Who this course is forThis course is designed for biologists, genomic variant analysts, clinical geneticists, and researchers working with short-read NGS data who want to strengthen their understanding of structural variants (CNVs). It is not a bioinformatics course and does not teach how to process raw NGS data. Instead, it focuses on the next stage of sequencing data analysis: evaluating whether a CNV is real, visualizing it in IGV, interpreting the type of rearrangement, and understanding which genes are affected. No advanced programming or bioinformatics expertise is required, making it accessible to professionals with basic knowledge of genetics and genomics.
Recommend Download Link Hight Speed | Please Say Thanks Keep Topic Live
No Password - Links are Interchangeable

